3.5.1.14: N-acyl-aliphatic-L-amino acid amidohydrolase
This is an abbreviated version!
For detailed information about N-acyl-aliphatic-L-amino acid amidohydrolase, go to the full flat file.
Reaction
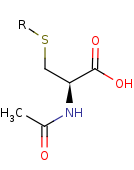 an N-acetyl-L-cysteine-S-conjugate +
an N-acetyl-L-cysteine-S-conjugate +
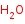 H2O=
H2O=
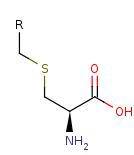 an L-cysteine-S-conjugate +
an L-cysteine-S-conjugate +
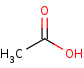 acetate
acetate
Synonyms
AA1, AA3, AAIII, ACY 1a, ACY 1b, ACY-1, ACY-1A, Acy1, acylase, acylase 1, acylase B, acylase I, alpha-N-acylaminoacid hydrolase, amido acid deacylase, aminoacylase, aminoacylase 1, aminoacylase 1a, aminoacylase 3, aminoacylase I, aminoacylase-1, aminoacylase-1A, aminoacylase-1B, benzamidase, D-AAse, D-Aminoacylase, dehydropeptidase II, DR_ACY, fatty acid-amino acid conjugate hydrolyzing aminoacylase 1, fMDF, fungus aminoacylase-1, hAcy1, hippurase, hippuricase, histozyme, HKA, hog intestinal acylase I, hog kidney aminoacylase, hog kidney aminoacylase I, intestine acylase I, L-ACY-1, L-amido-acid acylase, L-aminoacylase, LAA, lepidopteran aminoacylase 1, long acyl amidoacylase, M20, More, N-Acyl-D-aspartate amidohydrolase, N-acyl-L-amino acid amidohydrolase, N-acyl-L-amino acid hydrolase, N-acyl-L-amino-acid amidohydrolase, N-acyl-L-aminoacylase 1, N-acylase IA, N-formylmethionine deformylase, Nalpha-acyl-L-amino acid amidohydrolase, pAcy1, PhoACY, pig kidney aminoacylase, porcine kidney aminoacylase, short acyl amidoacylase, sphingosine kinase 1-interacting protein
ECTree
Subunits
Subunits on EC 3.5.1.14 - N-acyl-aliphatic-L-amino acid amidohydrolase
Please wait a moment until all data is loaded. This message will disappear when all data is loaded.
Please wait a moment until the data is sorted. This message will disappear when the data is sorted.
?

-
x * 42000, SDS-PAGE
?
-
x * 42000, SDS-PAGE
-
?
x * 64000, recombinant protein about 50000, SDS-PAGE
?
-
x * 43000, SDS-PAGE
-
dimer

-
2 * 40000, SDS-PAGE
dimer
-
2 * 43000, SDS-PAGE
dimer
-
2 * 36600, SDS-PAGE
dimer
-
2 * 40000, SDS-PAGE
dimer
-
2 * 42000, SDS-PAGE
dimer
-
2 * 44147, deduced from nucleotide sequence
dimer
-
2 * 42000, SDS-PAGE
-
dimer
-
2 * 44147, deduced from nucleotide sequence
-
dimer
-
1 * 26000 + 1 * 57000, SDS-PAGE
dimer
-
2 * 43000, SDS-PAGE
dimer
-
2 * 50000, native enzyme SDS-PAGE
dimer
-
1 * 61000 + 1 * 19000, SDS-PAGE, the enzyme is active only as complete heterodimer
dimer
-
2 * 43000, SDS-PAGE
dimer
-
dissociable enzyme, monomer and dimer forms
dimer
-
2 * 45000, denaturated intestinal mucosa enzyme, SDS-PAGE
dimer
2 * 45216, gel filtration and calculated from DNA sequence
dimer
2 * 45260, gel filtration and ESI-MS following a desalting step on a RP-HPLC column
homodimer

-
-
homodimer
2 * 45000 Da, SDS-PAGE
homotetramer

PAGE
homotetramer
-
4 * 43000, SDS-PAGE
homotetramer
-
SDS-PAGE before and after boiling with beta-mercaptoethanol
homotetramer
-
4 * 43000, SDS-PAGE, sedimentation studies
homotetramer
-
4 * 43814, calculated from sequence
homotetramer
-
4 * 43815, calculated from sequence
monomer

-
1 * 54000, SDS-PAGE
monomer
-
1 * 45000, SDS-PAGE
monomer
-
1 * 44000, denaturing and reducing gels
monomer
1 * 55000, purified wild type enzyme, SDS-PAGE
tetramer

-
-
tetramer
-
4 * 42000, SDS-PAGE
tetramer
-
4 * 43000, 4gatom Zn per mol enzyme, SDS-PAGE
tetramer
4* 42000 SDS-PAGE, recombinant enzyme
trimer

-
3 * 44000, gel filtration and SDS-PAGE
trimer
-
3 * 45847, deduced from deduced aminacid sequence
additional information

-
dimerization domain residues play important roles in binding and catalysis, overview
additional information
structure molecular modelling
additional information
structural homology modeling, overview
additional information
-
structural homology modeling, overview




 results (
results ( results (
results ( top
top






