5.5.1.4: inositol-3-phosphate synthase
This is an abbreviated version!
For detailed information about inositol-3-phosphate synthase, go to the full flat file.
Word Map on EC 5.5.1.4 
Reaction
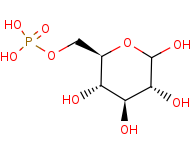 D-glucose 6-phosphate=
D-glucose 6-phosphate=
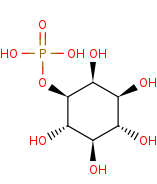 1D-myo-inositol 3-phosphate
1D-myo-inositol 3-phosphate
Synonyms
1D-myo-inositol 3-phosphate synthase, 1L-myo-Inositol-1-phosphate synthase, AtMIPS1, AtMIPS2-1, AtMIPS3, CaMIPS1, CaMIPS2, D-Glucose 6-phosphate cycloaldolase, D-Glucose 6-phosphate-1L-myoinositol 1-phosphate cyclase, D-Glucose 6-phosphate-1L-myoinositol 1-phosphate cycloaldolase, D-Glucose 6-phosphate-L-myo-inositol 1-phosphate cyclase, D-Glucose-6-phosphate,L-myo-inositol-1-phosphate cycloaldolase, D-myo-inositol-3-phosphate synthase, Glucocycloaldolase, Glucose 6-phosphate cyclase, Glucose-6-phosphate inositol monophosphate cycloaldolase, INO-1, INO-2, INO1, Ino1p, iNOS, Inositol 1-phosphate synthase, Inositol 1-phosphate synthetase, inositol 3-phosphate synthase, inositol phosphate synthase, inositol-1-phosphate synthase, inositol-3-phosphate synthase, INPS, INPS1, INS (3) P1 synthase, IP synthase, IPS, L-myo-inositol 1-phosphate synthase, L-myo-Inositol 1-phosphate synthetase, L-myo-Inositol-1-phosphate synthase, MI-1-P synthase, MIP synthase, MIPS, MIPS1, MIPS2, myo-D-inositol 3-phosphate synthase, myo-inositol 1-phosphate synthase, myo-inositol 3-phosphate synthase, myo-inositol phosphate synthase, myo-Inositol-1-P synthase, Myo-inositol-1-phosphate synthase, myo-inositol-3-phosphate synthase, OsMIPS1, OsMIPS1-1, OsMIPS1-2, OsMIPS1-3, OsMIPS2, PcINO1, PINO1 protein, PIP synthase, PIPS, RINO1 protein, SaINO1, sll1981, Synthase, myo-inositol 1-phosphate, TaMIPS1, TaMIPS2, tbINO, yMIP synthase, [Ins(3)P1] synthase
ECTree
KM Value
KM Value on EC 5.5.1.4 - inositol-3-phosphate synthase
Please wait a moment until all data is loaded. This message will disappear when all data is loaded.
Please wait a moment until the data is sorted. This message will disappear when the data is sorted.
1.15
2-deoxy-D-glucose 6-phosphate
pH 7, 37°C, Vmax: 280 nM
0.12 - 4.47
D-glucose 6-phosphate
0.035 - 2.85
glucose 6-phosphate
additional information
additional information
-
0.12
D-glucose 6-phosphate

-
90°C, pH 7.5
0.24
D-glucose 6-phosphate
-
with addition of 5% ethanol plus 50 mM KCl
0.26
D-glucose 6-phosphate
-
NAD+, chloroplastic enzyme
0.26
D-glucose 6-phosphate
-
with addition of 20 mM NH4Cl
0.67 - 0.72
D-glucose 6-phosphate
-
pH 7.5, 30°C, dependent on the assay method, presence or absence of Mg2+
0.8
D-glucose 6-phosphate
-
pH 7.6, 37°C
0.9
D-glucose 6-phosphate
-
with addition of 5% ethanol
0.922
D-glucose 6-phosphate
Gleichenia glauca
-
pH 7.5
0.97
D-glucose 6-phosphate
-
pH 7.5, recombinant refolded enzyme
1.05
D-glucose 6-phosphate
-
pH 7.0-7.5, 30°C
1.18
D-glucose 6-phosphate
-
-
1.19
D-glucose 6-phosphate
-
pH 7.5, native enzyme
1.2
D-glucose 6-phosphate
-
-
1.8
D-glucose 6-phosphate
native enzyme
1.9
D-glucose 6-phosphate
-
-
1.97
D-glucose 6-phosphate
native enzyme
2.5
D-glucose 6-phosphate
wild-type
2.5
D-glucose 6-phosphate
recombinant enzyme
2.63
D-glucose 6-phosphate
isozyme MIPS1
2.7
D-glucose 6-phosphate
-
-
2.7
D-glucose 6-phosphate
isozyme MIPS2
2.83
D-glucose 6-phosphate
mutant lacking amino acids 174-210
3
D-glucose 6-phosphate
recombinant enzyme
3
D-glucose 6-phosphate
wild-type, 37°C
3.19
D-glucose 6-phosphate
mutant bearing amino acids 174-210 of Porteresia coarctata enzyme, 37°C
3.4
D-glucose 6-phosphate
-
presence of 0.6 mM valnoctamide, pH 7.6, 37°C
3.5
D-glucose 6-phosphate
-
pH 7.5, 37°C, 7.5, recombinant enzyme
3.89
D-glucose 6-phosphate
-
-
4.4
D-glucose 6-phosphate
-
-
4.47
D-glucose 6-phosphate
-
-
0.035
glucose 6-phosphate

-
-
0.06
glucose 6-phosphate
-
-
0.33
glucose 6-phosphate
-
-
0.5
glucose 6-phosphate
-
-
0.52
glucose 6-phosphate
-
-
0.57
glucose 6-phosphate
-
-
1.95
glucose 6-phosphate
-
cytosolic enzyme from streptomycin-bleached cells, and chloroplastidic enzyme
1.97
glucose 6-phosphate
-
cytosolic enzyme
2.14
glucose 6-phosphate
-
chloroplastic enzyme
2.17
glucose 6-phosphate
-
cytosolic enzyme
2.25
glucose 6-phosphate
-
cytosolic enzyme from green cells
2.51
glucose 6-phosphate
-
cytosolic enzyme from dark-grown cells
2.85
glucose 6-phosphate
-
chloroplastic enzyme
0.0051
NAD+

-
90°C, pH 7.5
0.08
NAD+
-
chloroplastic enzyme
0.11
NAD+
-
cytosolic enzyme
0.12
NAD+
-
chloroplastic enzyme
0.14
NAD+
-
cytosolic enzyme
0.16
NAD+
-
cytosolic enzyme from green cells
0.19
NAD+
-
cytosolic enzyme from dark-grown cells
0.2
NAD+
-
cytosolic enzyme from streptomycin-bleached cells
0.42
NAD+
-
pH 7.5, 37°C, 7.5, recombinant enzyme
0.9
NAD+
Gleichenia glauca
-
pH 7.5
1.67
NAD+
-
pH 7.5, recombinant refolded enzyme
1.95
NAD+
-
pH 7.5, native enzyme
additional information
additional information

-
-
-
additional information
additional information
-
-
-
additional information
additional information
-
-
-
additional information
additional information
-
-
-
additional information
additional information
-
-
-
additional information
additional information
-
-
-
additional information
additional information
-
-
-
additional information
additional information
-
-
-
additional information
additional information
-
-
-
additional information
additional information
-
-
-
additional information
additional information
-
-
-
additional information
additional information
-
-
-
additional information
additional information
-
dissociation constants
-
additional information
additional information
-
Vmax is 2.95 mM
-
additional information
additional information
-
kinetics, effects of different divalent metal ions, overview
-




 results (
results ( results (
results ( top
top






