3.6.5.5: dynamin GTPase
This is an abbreviated version!
For detailed information about dynamin GTPase, go to the full flat file.
Reaction
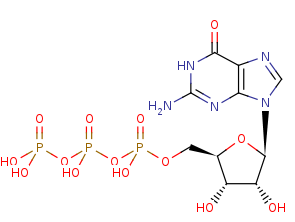 GTP +
GTP +
 H2O=
H2O=
 GDP +
GDP +
 phosphate
phosphate
Synonyms
AtDRP1A, B-dynamin , caSey1p, D100 , dDyn, DLC1, DLP-1, DLP1, DlpA, DlpB, DlpC, Dnm1, DNM1L, DNM2, DNM3, DRP, Drp1, Drp1-long, Drp1-short, DRP1E, DRP5A, Drp6, DrpB, Dyn, Dyn I, Dyn-1, Dyn1, DYN2, Dyn2aa, Dyn2ab, Dyn2ba, Dyn2bb, Dyn3, dynamin, dynamin 1, dynamin 1 GTPase, dynamin 1aa, dynamin 2, dynamin 2aa, dynamin 3, dynamin 3aa, Dynamin BREDNM19 , dynamin family GTPase, dynamin GTPase, dynamin I, dynamin I GTPase, dynamin I guanosine triphosphatase, Dynamin UDNM , Dynamin, brain , Dynamin, testicular , dynamin-1, dynamin-1 GTPase, dynamin-1-like GTPase, dynamin-1-like protein, dynamin-2, dynamin-3, dynamin-like GTPase, dynamin-like guanosine triphosphatase, dynamin-like guanosine-triphosphate hydrolase, dynamin-like myxovirus resistance protein A, dynamin-like protein 1, dynamin-like protein 6, dynamin-related GTPase, dynamin-related protein, dynamin-related protein 1, dynamin-related protein 1 isoform 3, dynamin-related protein 1A, dynamin-related protein 1E, dynamin-related protein 2A, dynamin-related protein B, dynamin-retated GTPase Drp1, dynamin1, dynamin2, dynamin2 GTPase, dynamine-related GTPase, dynein light chain 1, dynI, dynI GTPase, EC 3.6.1.50, fission dynamin, GTP phosphohydrolase, GTPase, GTPase dynamin, GTPase dynamin-1, GTPase dynamin-related protein 1, guanine triphosphatase, guanosine 5'-triphosphatase, guanosine triphosphatase, hDrp1 isoform 3, large GTPase, Mfn1, mitochondrial division dynamin, mitochondrial dynamin, mitochondrial inner membrane fusion dynamin Mgm1, mitofusin 1, mOPA1, multidomain GTPase, Mx GTPase, MX1, MxA, myxovirus resistance GTPase, neurolastin, Nicotiana tabacum dynamin-related protein 3, NtDRP3, OPA1, optic atrophy 1, PfDYN2, phosphatase, guanosine tri-, Protein shibire, ribosomal GTPase, Sey1p, Shibire protein , T-dynamin , Vps1p
ECTree
KM Value
KM Value on EC 3.6.5.5 - dynamin GTPase
Please wait a moment until all data is loaded. This message will disappear when all data is loaded.
Please wait a moment until the data is sorted. This message will disappear when the data is sorted.
additional information
additional information
-
0.0034
GTP

-
22°C, wild-type dynamin, basal GTPase activity
0.005 - 0.05
GTP
-
Km is in the range of 0.005-0.05 mM depending on salt and temperature conditions
0.0053
GTP
-
22°C, wild-type dynamin, lipid tubule stimulated GTPase activity
0.0067
GTP
-
K142A mutant of the GTPase domain of dynamin
0.0076
GTP
-
T141Q mutant of the GTPase domain of dynamin
0.0078
GTP
-
wild-type dynamin
0.0091
GTP
-
T65A mutant of the GTPase domain of dynamin
0.025
GTP
-
in presence of microtubules
0.025
GTP
-
I690K mutant, high salt
0.025
GTP
-
I690K mutant, low salt
0.025
GTP
-
K694A mutant, low salt
0.0253
GTP
-
in 10 mM Tris-HCl, 10 mM NaCl, 2 mM Mg2+, at 30°C
0.028
GTP
-
I690K mutant, + GAP
0.031
GTP
-
wild-type dynamin, + GAP
0.033
GTP
-
K694A mutant, + GAP
0.035
GTP
-
wild-type dynamin, low salt
0.037
GTP
-
37°C, wild-type dynamin, lipid tubule stimulated GTPase activity
0.04
GTP
-
wild-type dynamin, high salt
0.041
GTP
-
37°C, S61A mutant dynamin: lipid tubule stimulated GTPase activity, T65D mutant dynamin: basal GTPase activity
0.044
GTP
-
K694A mutant, high salt
0.046
GTP
-
37°C, wild-type enzyme
0.056
GTP
-
37°C, S61D and T141A mutant dynamin, lipid tubule stimulated GTPase activity
0.065
GTP
-
R66A mutant of the GTPase domain of dynamin
0.067
GTP
-
37°C, S61A mutant dynamin, basal GTPase activity
0.068
GTP
-
37°C, S61D mutant dynamin, basal GTPase activity
0.096
GTP
-
37°C, mutant enzyme R59K
0.102
GTP
-
37°C, wild-type dynamin, basal GTPase activity
0.149
GTP
-
37°C, T65H mutant dynamin, basal GTPase activity
0.174
GTP
-
37°C, T141A mutant dynamin, basal GTPase activity
0.241
GTP
-
37°C, mutant enzyme R59A
0.37
GTP
-
in absence of microtubules
0.513
GTP
-
37°C, T65D mutant dynamin, lipid tubule stimulated GTPase activity
0.763
GTP
-
37°C, T141D mutant dynamin, basal GTPase activity
0.88
GTP
-
37°C, T65A mutant dynamin, basal GTPase activity
0.988
GTP
-
37°C, T65H mutant dynamin, lipid tubule stimulated GTPase activity
1.94
GTP
-
37°C, T141D mutant dynamin, lipid tubule stimulated GTPase activity
2.115
GTP
-
37°C, T65A mutant dynamin, lipid tubule stimulated GTPase activity
additional information
additional information

Michaelis-Menten kinetics
-
additional information
additional information
Michaelis-Menten kinetics
-
additional information
additional information
-
kinetic data, temperature-dependent effects on kinetic parameters, self-assembly alters the Km for GTP
-
additional information
additional information
steady-state GTPase activities of recombinant Drp1 GTPase-GTPase effector fusion protein
-
additional information
additional information
-
steady-state GTPase activities of recombinant Drp1 GTPase-GTPase effector fusion protein
-




 results (
results ( results (
results ( top
top






